
4A Data
Management: Acquiring, Acting-on, Archiving and Advertising data at the
University of Western Sydney by Peter
Sefton and Peter Bugeia is licensed under a Creative Commons
Attribution 3.0 Unported
License.
Slide 1

Notes
Abstract
There has been significant Government investment in Australia in repository and eResearch infrastructure over the last several years, to provide all universities with an institutional repository for publications, and via the Australian National Data Service to encourage the creation of institution-wide Research Data Catalogues, and research Data Capture applications. Further rounds of funding have added physical data storage and cloud computing services. This presentation looks at an example of how these streams of money have been channeled together at the University of Western Sydney to create a joined-up vision for research data management across the institution and beyond, creating an environment where data may be used by research teams within and outside of the institution. Alongside of the technical services, we report on early work with researchers to create a culture of replicable use of data, towards the vision of truly reproducible research.
This presentation will show a proven end-to-end design for research data flows, starting from a research group, The Hawkesbury Institute for the Environment, where a large sensor network gathers data for use by institute researchers, in-situ, with data flowing-through to an institutional data repository and catalogue, and thence to Research Data Australia - a national data search engine. We also discuss a parallel workflow with a more generic focus - available to any researcher. We also report on work we have done to improve metadata capture at source, and to create infrastructure that will support the entire research data lifecycle. We include demonstrations of two innovations which have emerged from the associated project work: the first is of a new tool for researchers to find, organize, package and publish datasets; the second is of a new packaging format which has both human-readable and machine-readable components.
Slide 2

Notes
Some of the work we discuss here was funded by the Australian National Data Service. See:
Seeding the commons project to describe data sets at UWS and the Data catalogue project.
HIEv Data Capture at the Hawkesbury Institute for the Environment
The talk
Notes
We'll use the four A's to talk about some issues in data management.
-
We need a simple framework which covers it all, to capture how we work with research data from cradle to grave:
-
We need to Acquire the raw data and make it secure and available to be worked on.
-
We need to Act on the data to cleanse it while keeping track of how it was cleansed, analyse it using tools to support our research, while maintaining the data's provenence.
-
We need to Archive the data from working storage to an archival store, making it citable
-
We need to Advertise that the data exists so that others can discover it and use it confidently with simple access mechanisms and simple tools.
-
4A must work for
-
high-intensity research data such as that from gene sequences, sensor networks, astronomy, medical diagnostic equipment, etc.
-
the long tail of unstructured research data.
For example
Notes
In the presentation, I used a short video on how to catch a kangaroo. (Late the night before I was searching for this video and I forgot how spell kagaroo and tried starting it like this “How to catch a C … A ... N ...” - at which point the Google suggestion popped up with this, which I decided not to show at the conference I'd blame the jet-lag, but you wouldn't believe me.)
If only data capture were as simple as catching a kangaroo in a shopping bag!
Australian Government Initiatives in Research Data Management

Notes
There have been several rounds of investment in (e)research infrastructure in Australia over the last decade, including substantial investments to get institutional publications repositories established.
-
Australian National Data Service (ANDS) $50M (link)
-
National eResearch Collaboration Tools and Resources (NeCTAR) project (link) $50M
-
Research Data Storage Infrastructure (RDSI) $50M (link)
-
Implemented to date:
-
National Research Data Catalogue - Research Data Australia
-
Standard approach to updating the Catalogue (OAI-PMH and rif-cs)
-
10+ Institutional Metadata Repositories implemented
-
120+ data capture applications implemented across 30+ research organisations
-
Upgrade of High Performance Computing infrastructure
-
Colocation of data storage and computing
Slide 6

Notes
UWS is a young ( ~20years) university performing well above most of its contemporaries in research.
Slide 7
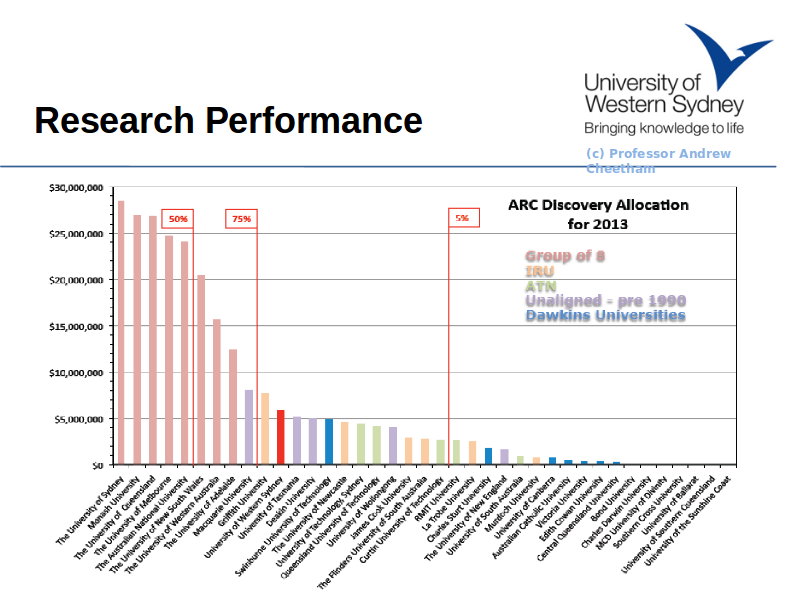
Notes
This slide by Prof Andrew Cheetham - the Deputy Vice Chancellor for Research shows that UWS performs very well at attracting competitive grant income from the Australian Research Council.
Slide 8

Notes
UWS is concentrating its research into flagship institutes - we will be talking in more detail about HIE, here, our environmental institute which does research from cutting across different disciplines spanning from the leaf level to the ecosystem level.
Slide 9

Notes
Slide 11

Notes
These are Intersect's members. Intersect also collaborates with other eResearch organisations throughout Australia.
The slide is a photo of at the recent Hackfest event. This is an annual fun competition for software developers to use open government data in innovative ways. Intersect hosted the NSW chapter of the event.
eResearch @ UWS

Notes
The eResearch unit at UWS is a small team, currently reporting to the Deputy Vice Chancellor, Research. See our FAQ.
Slide 13
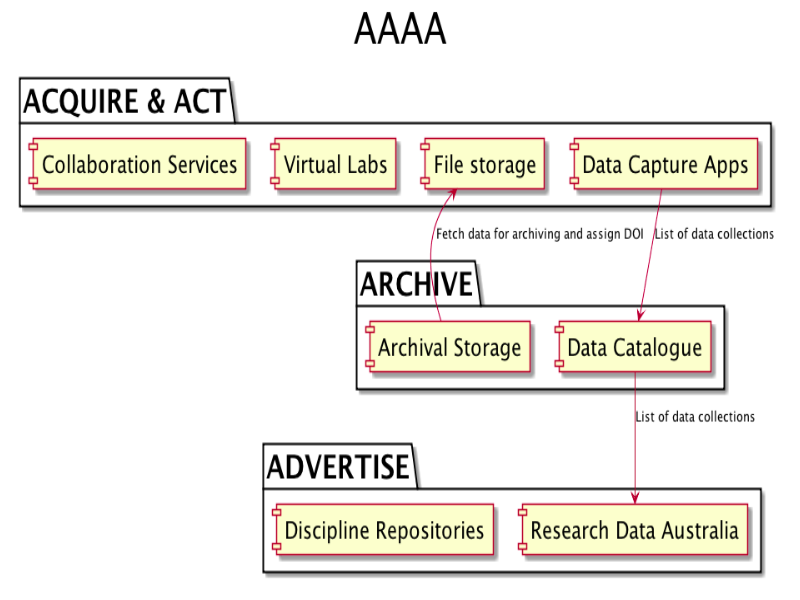
Notes
At UWS, we haven't tried to drive change with top-down policy. Instead, we've taken a practical, project-based approach which has allowed a data architecture to evolve. The eResearch Roadmap calls for a series of data capture applications to be developed for data-intensive research, along with a generic application to cover the long tail of research data.
The 4A Vision
For the purposes of this presentation we will talk about the ‘4A’ approach to research data management - Acquire, Act, Archive and Advertise. The choice of different terms from the 2Rs Reuse and Reproduce of the conference theme is intended to throw a slightly different light on the same set of issues. The presentation will examine each of these ‘A’s in turn and explain how they have helped us to organize our thinking in developing a target technical data architecture and integrated data-related end-to-end business processes and services involving research technicians and support staff, researchers and their collaborators, library staff, information technology staff, office of research services, and external service providers such as the Australian National Data Service and the National Library of Australia. The presentation will also discuss how all of this relates to the research project life cycle and grant funding approval.
Acquiring the data
We are attacking data acquisition (known as Data Capture by the Australian National Data Service, ANDS 1) in two ways:
With discipline specific applications for key research groups. A number of these have been developed in Australia recently (for example MyTARDIS 2), we will talk about one developed at UWS. With ANDS funding, UWS is building an open source automated research data capture system (the HIEv) for the Hawkesbury Institute for the Environment to automatically gather time-series sensor data and other data from a number of field facilities and experiments, providing researchers and their authorised collaborators with easy self-service discovery and access to that data.
Generic services for Data storage via simple file shares, Integration with cloud storage including Dropbox.com and other distributed file systems. And Source-code repositories such as public and private github and bitbucket stores for working code and textual data.
Acting on data
The data Acquisition services described above are there in the first instance to allow researchers to use data. With our environmental researchers, we are developing techniques for developing reusable data sets which include raw data, commented scripts to clean the data (eg a comment “filter out known bad-days when the facility was not operating”) then re-organize it via resampling or other operations into useful ‘clean’ data that can be fed to models, plotted etc and used as the basis of publications. Demo: the presentation will include a live demonstration of using HIEv to work on data and create a data archive.
From action to archive
Having created both re-usable base data sets and publication-specific operations on data to create plots etc there are several workflows where various parties trigger deposit of finished, fixed, citable data into a repository. Our project team mapped out several scenarios where data are deposited with different actors and drivers including motivations that are both carrot (my data set will be cited) and stick (the funder/journal says I have to deposit). Services are being crafted to fit in with these identified workflows rather than build new things and assume “they will come”.
Archiving the data
The University of Western Sydney has established a Research Data Repositoryi (RDR), the central component of which is a Research Data Catalogue, running on the ReDBOX open source repository platform. While individual data acquisition applications such as HIEv are considered to have a finite lifespan, the RDR will provide on-going curation of important research datasets. This service is set up to harvest data sets from the working-data applications, including the HIEv data-acquisition application and the CrateIt data packaging service using the Open Archives Initiative – Protocol for Metadata Harvesting (OAI-PMH).
Advertising the data
As with Institutional Publications Repositories, one of the key functions of the Research Data Repository is to disseminate metadata about holdings to aggregation services and give data a web presence. Many Australian institutions are connected to the Research Data Australia discovery service 6, which harvests metadata via an ANDS-defined standard over the OAI-PMH harvesting protocol. There is so far no Google-Scholar-like service which is harvesting data about data sets via direct web crawling (that we know about), so there are no firm standards for how to embed data in a page, but we are tracking the developments of the Schema.org vocabulary, which is driven largely by Google’s group of companies which are Google’s peers, and the work described above on data packaging with RDFa metadata is intended to be consumed by direct crawlers. It is possible to unzip a CrateIt package and expose it to the web thus creating a machine-readable entry-point to the data within the Zip/BagIt archive.
Looking to the future, the University is also considering plans for an over-arching discovery hub, which would bring together all metadata data about research including information on publications, people, and organisation.
Technical architecture
The following diagram shows the first end-to-end data capture to archiving pathways to be turned on at the University of Western Sydney, covering Acquisition and Action on data (use) and Archiving and Advertising of data for reuse. Note the inclusion of a name-authority service which is used to ensure that all metadata flowing through the system is unambiguous and inked-data-ready 7. The name Authority is populated with data about people, grants and subject codes from databases within the research services section of the university and from community-maintained ontologies. A notable omission from the architecture is integration with the Institutional Publications Repository – we hope to be able to report on progress joining up that piece of the infrastructure via a Research Hub at Open Repositories 2014.
i Project materials refer to the repository as a project which includes both working and archival storage as well as some computing resources, drawing a line around ‘the repository’ that is larger than would be usual for a presentation at Open Repositories.
Slide 14
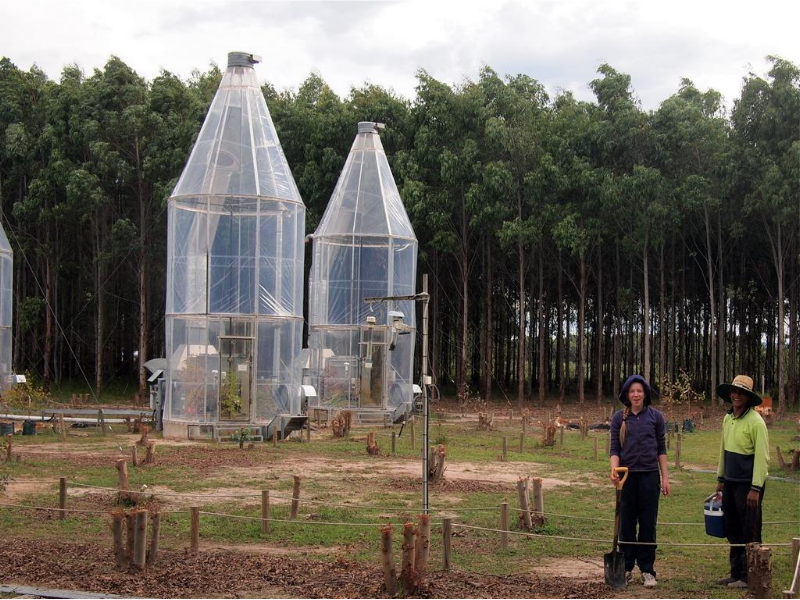

Notes
There are a number of major research facilities at HIE, here are two whole-tree chambers which allow control over temperature, moisture and atmospheric CO2.
Slide 15
Notes
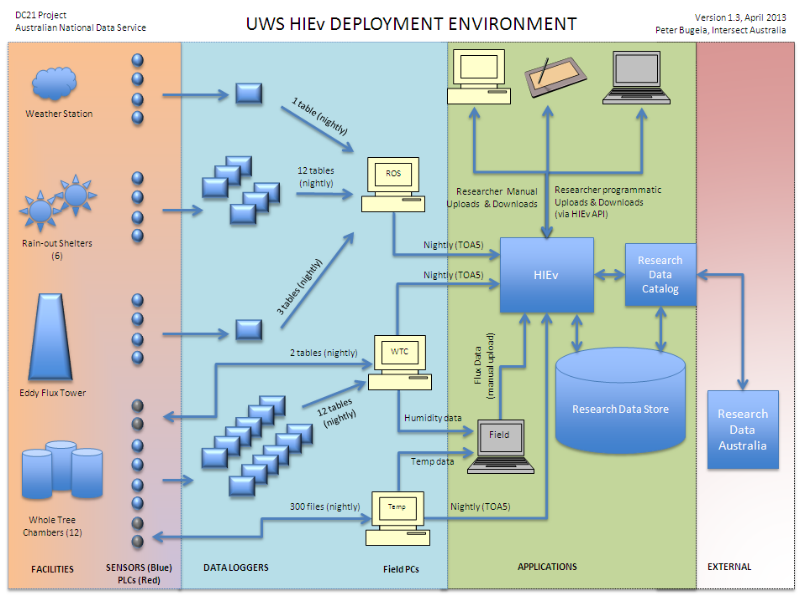
This diagram shows the end to end data and application architecture which Intersect and UWS eResearch built to capture data from HIE sensors and other sources. Each of the columns roughly equates to the four A model. Once data is packaged in the HIev, it is stored in the Research Data Store and there is a corresponding record for it in the Research Data Catalog. The data packaging format produced by the HIEv, along with the delivery protocol are key to the architecture: the data packaging format (based on bagit) is stand-alone from the HIEv and self-describing, the delivery protocol (OAI-PMH) is well-defined and standards based. THese are discussed in more detail in later slides. When other data capature applications are developed at UWS, to integrate into and extend the architecture they will simply need to package data in the same format and produce and deliver the same meta-data via the same delivery protocol as the HIEv.
Slide 16
Notes
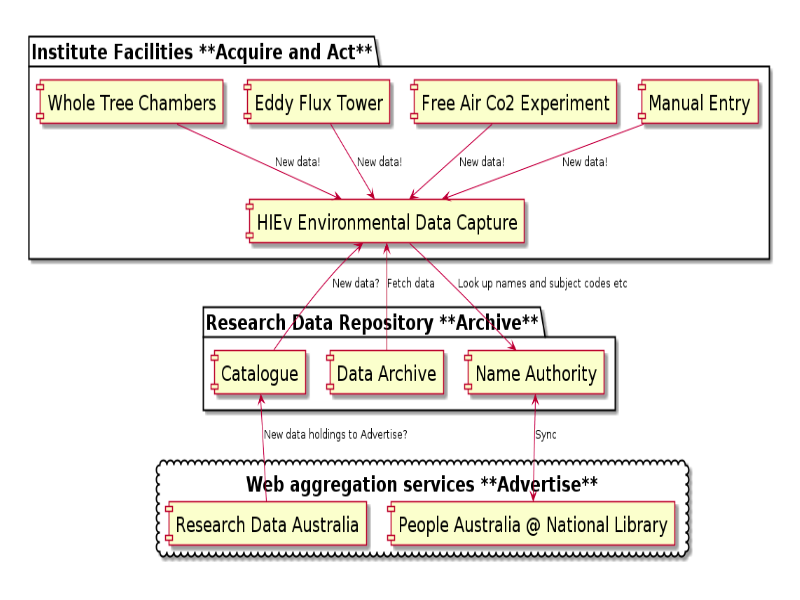
This diagram shows how the four 'A's fit together for HIE. Acquisition and action are closely related - it is important to provide services which researchers actually want to use and to build in data publishing and packaging services rather than setting up an archive, and hoping they come to it with data.
Slide 17
Notes
The HIEv/DC21 application is available as open source:
-
Funded by ANDS
-
Developed by Intersect
-
Automated data capture
-
Ruby on Rails application
-
Agile development methodology
-
Went live in Jan 2013.
-
1200 files, 15 GB of RAW data, 25 users.
-
120 files auto-uploaded nightly, +1GB per week
-
Expected to reach 50,000 files in next couple of years
-
Now extended to include Eucface data
-
Possibly to be extended to include Genomic data (20TB per year)
-
Integrated with UWS data architecture
-
Supports the full 4 As - links Acquire to Act to Archive
Slide 18
Notes
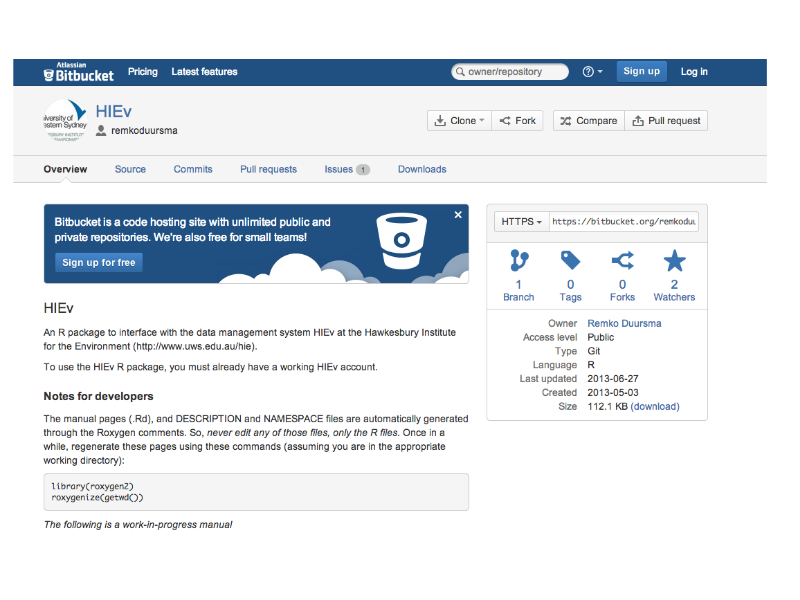
Acting on data: our researchers are not staring to do work with the HIEv system: here's an API developed by Dr Remko Duursma to consume data from R-stats.
Slide 19
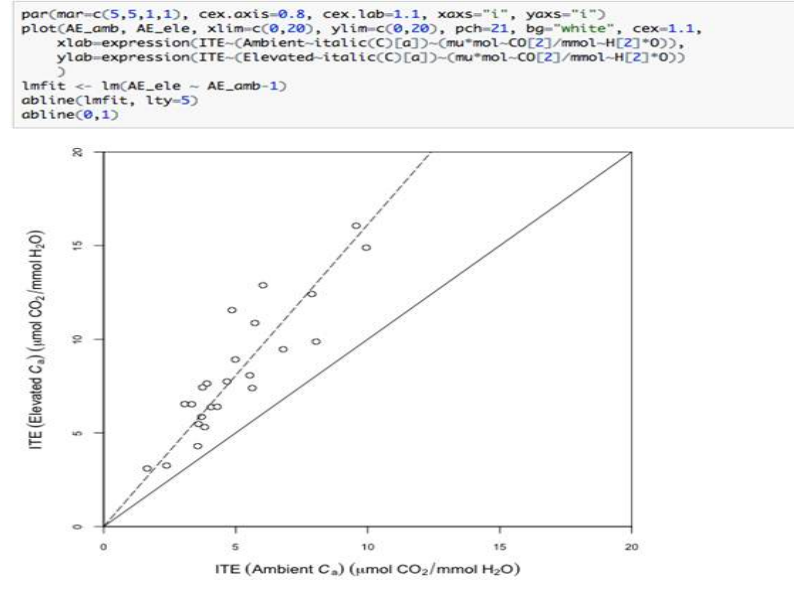
Notes
Acting on data: researchers can pull data either manually of via API calls and do work, such as this R-plot.
From acting to archiving...
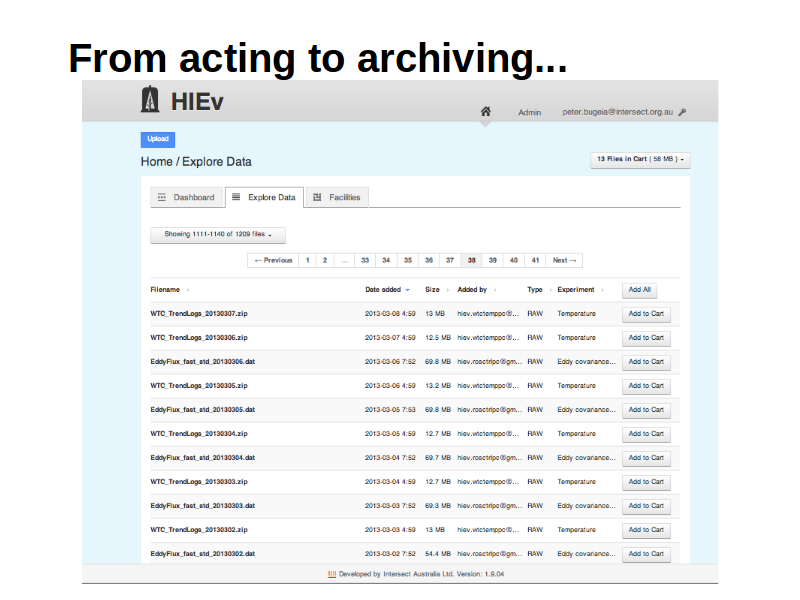
Notes
The following few slides show how a user can select some files...
Slide 21
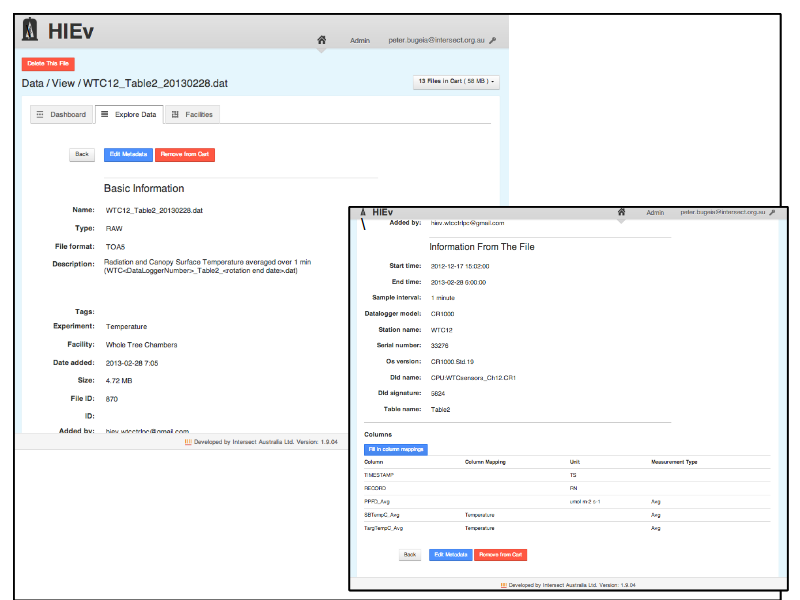
Notes
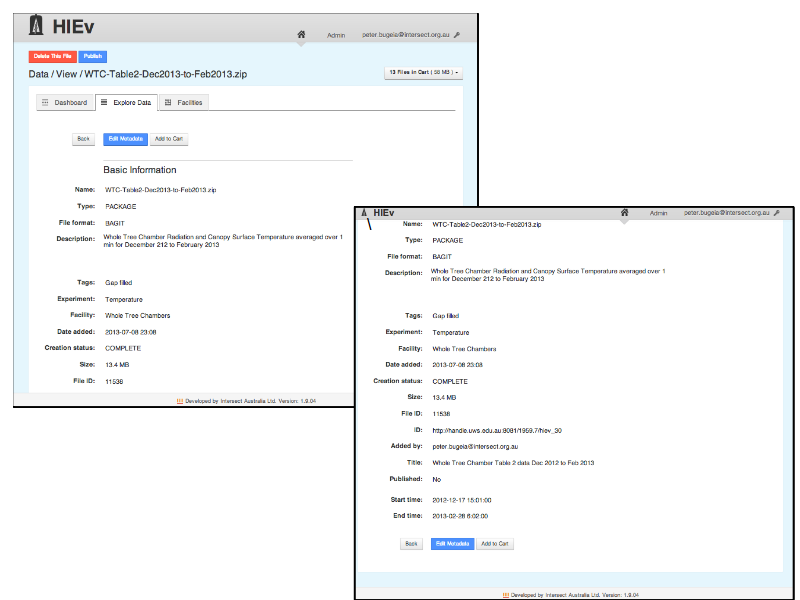
... look at file metadata ...
Slide 22
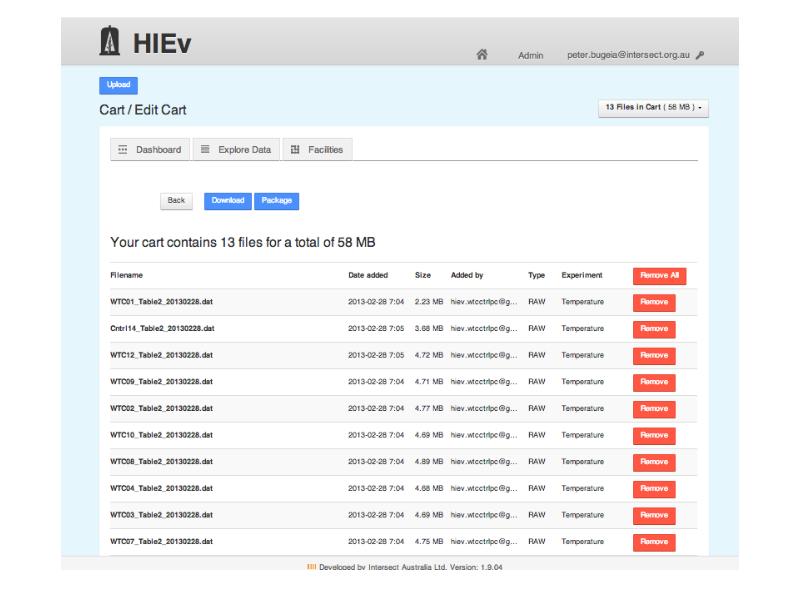
Notes
... add files to a cart ...
Slide 23
Notes
... download the files in a zip package ...
Slide 24
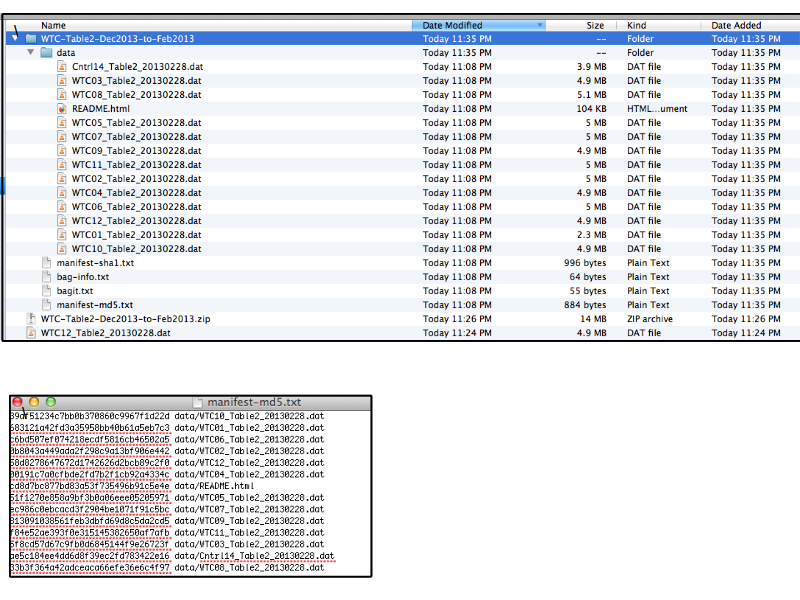
Notes
... inside the zip the files are structured using the bagit format ...
Slide 25
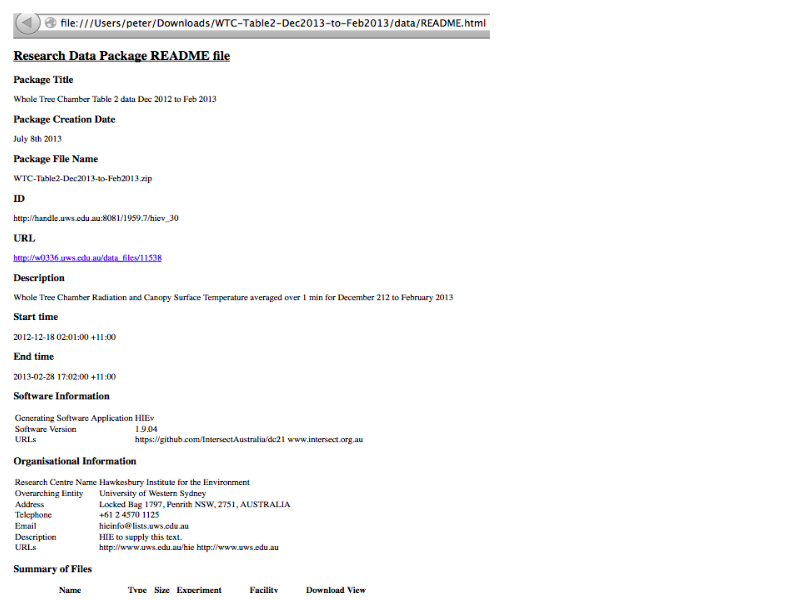
Notes
... with a standalone README.html file containing all the metadata we know about the files and associated research context (experiments, facilites) ...
Slide 26

Notes
... with detail about every file as per the HIEv application itself
Slide 27
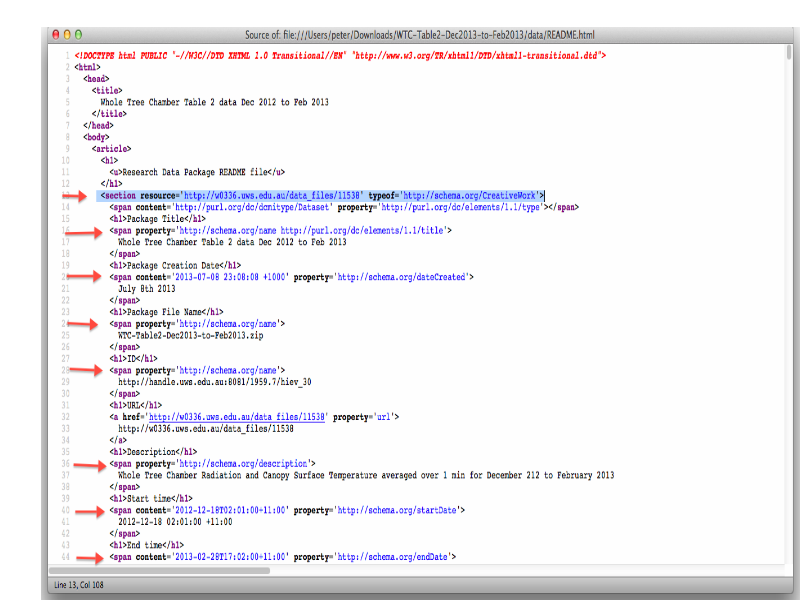
Notes
... and embedded machine readable metadata using RDFa lite attributes
Slide 28
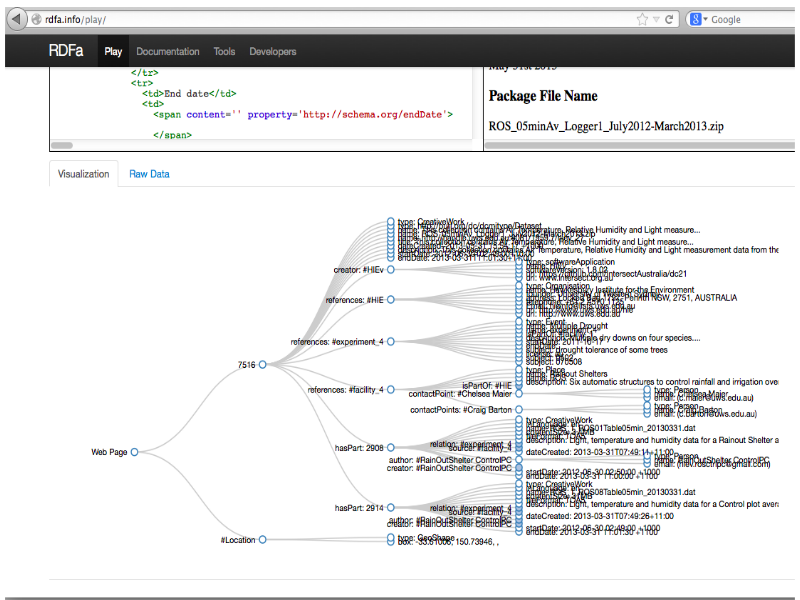
Notes
... the RDFa metadata describes the data-set as a graph.
Completed packages flow-through to the Research Data Catalogue via an OAI-PMH feed, and there they are given a DOI so they can be cited. The hand-off between systems is important, once a DOI is issued the data set has to be kept indefinitely and must not be changed.
Slide 29
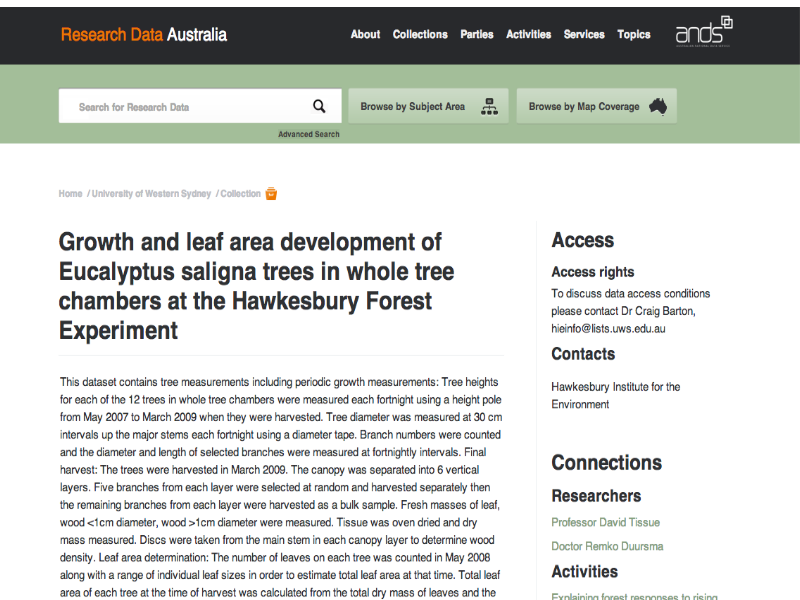
Notes
Advertising - data. This is a record about an experiment on Research Data Australia.
Slide 30

Notes
I said I'd talk about the long tail. He are two.
We looked in some detail at how the HIEv data capture application works for environmental data - but what about researchers who are on the long tail, and who don't have specific software applications for their group?
We are working on a similar Acquire and Act service that will operate with files and trying to make it as useful and attractive as possible. Most research teams we talk to at UWS are using Dropbox or one of the other 'Share, Sync, See' services. Dropbox has limitation on what we can do with its APIs and does not play nicely with authentication schemes other than its own, so we are looking at building 'Acquire and Act' services using an open source alternative; ownCloud.
Our application is known as Cr8it (Crate-it).
Slide 31
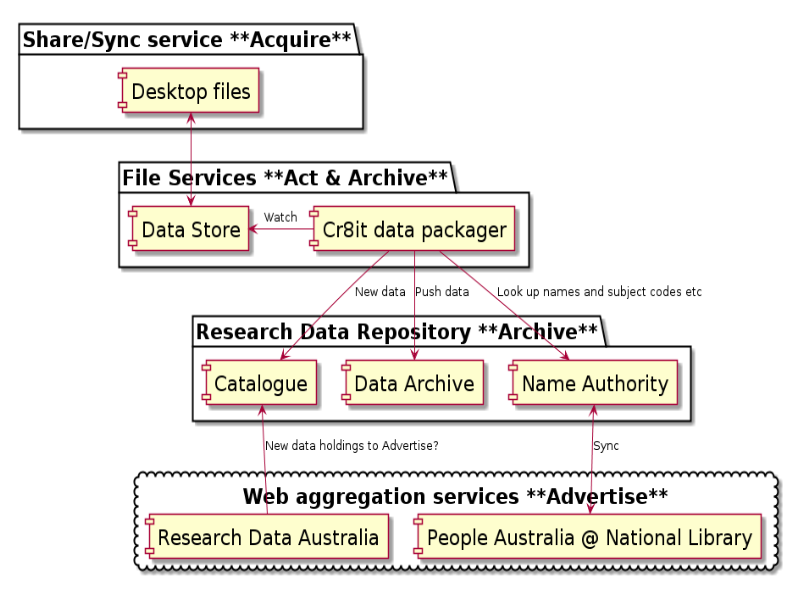
Notes
A number of techniques employed at UWS:
-
the "R" drive
-
research-project-oriented data shares
-
synchronisation with dropbox and owncloud
-
synchronisation with github and svn
References
1. Burton, A. & Treloar, A. Designing for Discovery and Re-Use: the ‘ANDS Data Sharing Verbs’ Approach to Service Decomposition. International Journal of Digital Curation 4, 44–56 (2009).
2. Androulakis, S. MyTARDIS and TARDIS: Managing the Lifecycle of Data from Generation to Publication. in eResearch Australasia 2010 (2010).at <http://ccaeducause1.caudit.edu.au/index.php/eraust/2010/paper/view/62>
3. Sefton, P. M. The Fascinator - Desktop eResearch and Flexible Portals. (2009).at <https://smartech.gatech.edu/handle/1853/28483>
4. Kunze, J., Boyko, A., Vargas, B., Madden, L. & Littman, J. The BagIt File Packaging Format (V0.97). at <http://tools.ietf.org/html/draft-kunze-bagit-06>
5. Group, W. W. & others RDFa Core 1.1 Recommendation. (2012).at <http://www.w3.org/TR/rdfa-syntax/>
6. Wolski, M., Richardson, J. & Rebollo, R. Shared benefits from exposing research data. in 32 nd Annual IATUL Conference (2011).at <http://iatul2011.bg.pw.edu.pl/proceedings/ft/Wolski_M.pdf>
7. Berners-Lee, T. Linked data, 2006. at <http://www.w3.org/DesignIssues/LinkedData.html>
\
\